Deciphering the protein signature of chronic pain
Pain is a major symptom of many diseases and the number one reason worldwide that people seek medical assistance. While acute pain serves protective functions, chronic pain presents a daunting challenge to patients and clinicians alike. In light of these difficulties, the identification of proteins specifically involved in chronic pain states would open new avenues for designing selective and efficient therapeutic interventions. Relevant studies are being conducted at the MPI for Experimental Medicine.
Yearbook article 2016, Max Planck Institute for Experimental Medicine, Göttingen
Emmy Noether Research Group Somatosensory Signal Transduction and Systems Biology
Author: Manuela Schmidt
Imagine …
… You encounter a good fairy who promises you that henceforth you can live free of pain. No doubt, you would agree to the fulfilment of this promise without batting an eyelid. But what would your everyday life be like without pain? You would get up in the morning, drink your coffee and fail to notice that it’s too hot and is scalding your tongue. On the sports field, you’d pay no heed to a knee injury and would play on, possibly causing permanent tissue damage. Because you’d feel no pain, you wouldn’t notice early enough that you have appendicitis with potentially life-threatening consequences. These are just a few examples that reflect the key protective function of pain as a warning signal. People whose sensation of pain is impaired by mutations in their genome provide particularly striking examples [1]. Their everyday life is marked by numerous, often serious injuries and inflammations, which often lead to premature death.
However, while such acute pain, known as nociceptive pain, acts as a vital warning sign of harmful conditions, chronic pain is a result of a maladaptation of the nervous system [2]. Chronic pain is a huge challenge, as it cannot be adequately treated with today’s pain medications. Patients with chronic pain are therefore exposed to considerable suffering, as well as serious treatment-related side effects. The latter are chiefly due to the fact that today's pain medications target proteins which occur throughout the body [3].
Therefore, to truly enjoy the benefits of a promised pain-free life, you would have to ask the good fairy to abolish only chronic pain. For this wish to be fulfilled, a better understanding of the unique molecular mechanisms that underlie chronic pain is essential.
Pain and membrane proteins
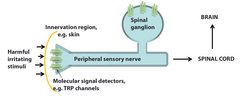
Pain is a major symptom of many diseases and the number one reason worldwide that people seek medical assistance. Vertebrates, including humans, have specialized somatosensory nerves whose cell bodies are grouped in the dorsal root ganglia parallel to the spinal cord. These somatosensory neurons innervate the skin and internal organs by means of thin cellular processes. To detect both normal and painful stimuli of varying quality (mechanical, thermal and chemical), the neurons carry proteins that act as primary molecular signal detectors. These membrane proteins (i.e. they are expressed in the plasma membrane of neurons) have the unique ability to detect and relay normal and harmful stimuli – from a gentle touch to the painful perception of a sharp nail [3].
It wasn’t until 15 to 20 years ago that pioneering research results deciphered the molecular identity of important signal detectors and significantly advanced research into the molecular basis of pain [3]. For example, several members of the transient receptor potential (TRP) ion channel family serve as a primary molecular signal detectors. These ion channels are activated by a variety of plant substances, such as capsaicin, the "hot" substance in fiery chilli peppers. The laboratory of David Julius (UCSF, USA) was able to show in mice that capsaicin activates a specific TRP ion channel (called TRPV1 channel) in sensory nerve fibres [4; 5]. This process triggers the fiery sensation that we associate with eating chilli peppers. Interestingly, the same ion channel is also activated by harmful heat, i.e. temperatures above 42°C, which also elicits a burning sensation [4; 5]. Another TRP channel, TRPA1, has been cloned and characterized by the laboratory of Ardem Patapoutian (Scripps Research Institute, USA). TRPA1 is activated by harmful cold (temperatures below 15°C), by a variety of irritating herbal and chemical substances, such as mustard oils, which are found in mustard and wasabi products [6], and by tear gases, which cause respiratory irritation [7].
These examples show that molecular sensors can detect an incredible repertoire of painful stimuli which can be either exogenous or endogenous (e.g. substances that are released during inflammation processes) in nature [7]. Activation of the signal detectors generates an electrical pulse that is converted by other membrane proteins into action potentials in the nerve. This action potential, in turn, triggers complex signal transduction cascades involving yet other membrane proteins in the spinal cord and ultimately in various brain regions, where the perception of pain is generated (Fig. 1).
The concept of “molecular machines”
Even this very brief description of the general principles of pain detection, transmission and perception illustrates the enormous importance of membrane proteins that are expressed in the “pain axis”, a term that refers to those regions of the peripheral nervous system (PNS) and central nervous system (CNS) that are involved in the processing of pain signals. Because the analysis of membrane proteins is fraught with technical difficulties, we know relatively little about which membrane proteins are relevant for specific types of pain and, in particular, details of how they are regulated. However, if it were possible to identify membrane proteins or regulatory mechanisms that are selectively active only during chronic pain, those processes could be used as targets for future therapies to render pain treatment more effective and reduce side effects [8].
However, research into the pathogenetic mechanisms of chronic pain has so far focussed mainly on the genomic or transcriptome level, i.e. the stage before translation of nucleic acids into proteins. Because proteins are the functional units of cells, and the transcriptome provides limited information on the proteome, i.e. the totality of proteins in a cell, a detailed proteomic analysis is essential. How is the activity of membrane proteins controlled during pain? To answer this question, it is important to identify associated or interacting proteins. At the heart of this approach is the concept of “molecular machines”, which states that the function of individual proteins can be dynamically modulated by their interaction with other proteins within multi-protein complexes [9]. Identifying the components of such protein complexes should consequently shed light on the function and regulation of proteins. The validity of this approach has been demonstrated in numerous examples. However, it has only stingingly been applied to the study of the molecular basis of pain.
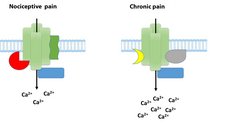
The declared goal of the Max Planck researchers is therefore to conduct comparative studies in order to examine the regulation of membrane proteins and associated protein networks during nociceptive and chronic pain. With this approach, the Research Group led by Manuela Schmidt has shown that an interacting protein, Annexin A2, reduces the activity of TRPA1 channels in mice during both nociceptive and chronic pain [10]. This work was honoured with the Förderpreis für Schmerzforschung der Deutschen Schmerzgesellschaft e. V. In the next step, the Max Planck researchers took on a much more difficult task: using advanced quantitative mass spectrometry, they searched for protein complexes that are associated with TRPA1 channels during chronic pain (this work was carried out in collaboration with Dr. Olaf Jahn, MPIem, and Dr. Henning Urlaub, MPIbpc). In fact, their findings suggest that TRPA1 channels form complexes with different proteins, depending on the applied pain paradigm, i.e. depending on whether nociceptive or chronic pain has previously been triggered in the mice (Fig. 2). The researchers also obtained similar results for the TRPV1 channels described above.
Prospects
Current projects of the Somatosensory Signalling and Systems Biology Research Group at the Max Planck Institute for Experimental Medicine are focussing on the characterization of novel, previously undescribed components of TRP channel-protein complexes and their pathological relevance to chronic pain in mice. In addition to the candidate-focused approach described here, i.e. studies that deal with specific TRP channels, the research group is also using a systems biology approach to investigate the differential regulation of several thousand proteins during various forms of pain. This work was awarded the German Pain Society’s Max von Frey Award in 2015. This progress was made possible by combining mouse models, biochemical work and the latest developments in mass spectrometry proteomic analysis. The expectation is that such studies will provide new insights into the molecular signature of pain at the protein level – knowledge that is essential for the development of efficient and targeted therapies.
Nature 389, 816-24 (1997)