Epigenetic changes don’t last
First comprehensive inventory of epigenetic changes over several generations shows that these often do not last and therefore probably have limited effects on long-term evolution
Jean-Baptiste Lamarck would have been delighted: geneticists no longer dismiss out of hand his belief that acquired traits can be passed on to offspring. When Darwin published his book on evolution, Lamarck’s theory of transformation went onto the ash heap of history. But in the last decade, we have learned that the environment can after all leave traces in the genomes of animals and plants, in form of so-called epigenetic modifications. Scientists at the Max Planck Institute of Developmental Biology in Germany have now produced the first comprehensive inventory of spontaneous epigenetic changes. Using Arabidopsis, the workhorse of modern plant genetics, the researchers determined how often and where in the genome epigenetic modifications occur – and how often they disappear again. They found that epigenetic changes are many orders of magnitude more frequent than conventional DNA mutations, but also often short lived. They are therefore probably much less important for long-term evolution than previously thought.
The team around Detlef Weigel, director of the Department for Molecular Biology, focused on one of the most important epigenetic marks, methylation of DNA. Tiny chemical building blocks, methyl groups, are thereby attached to individual letters of the DNA, mostly to cytosines. The genetic information itself in form of the four different letters or nucleotides that make up the genetic code remains unchanged in this process. To determine the rate and distribution of methylation changes in the genome, the German biologists looked at ten Arabidopsis lines. These lines came from the same stock, but had been propagated independently for 30 generations by self-fertilization. In the genome of the last generation the scientists then searched for differences in the methylation pattern in comparison to the common ancestor. They produced for each individual a complete map of methylated cytosines in the genome, the so-called methylome.
“For each line, we were able to look at about 14 million cytosines,” said Claude Becker, a member of the Tübingen team. On average, every plant had almost 3 million methylated cytosines. The vast majority of these were the same in all lines, but about 6 percent had changed since the lines had become separated. At these positions, at least one of the individuals was different, with either methylation gained or lost relative to the ancestor. Each of the lines had about 30,000 such epimutations, which was 1,000 times more than DNA mutations.
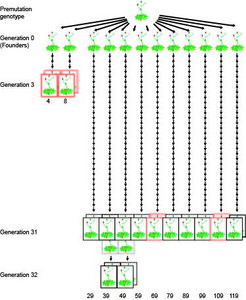
With 30,000 epimutations after 30 generations, the geneticists had expected that 1,000 epimutations occurred in each generation. When they directly compared parents and their immediate offspring, they were surprised to find that the epimutation rate was three to four times as high. The scientists concluded that many epimutations are apparently not stable and return to their original state after a few generations. Thus, averaging the mutation rate over many generations is misleading. Becker’s colleague Jörg Hagmann therefore cautions not to overestimate the importance of DNA methylation during evolution: “Our experiments show that methylation changes are often reversible”. In other words: New epimutations are often not maintained over the long term. “Only when selection wins out over reversion can these epimutations affect evolution,” says Hagmann. A new epimutation thus must have a strong evolutionary advantage so that it can become established before being lost again. Because reverse mutations do not necessarily happen in the next generation, it is still possible that epigenetic differences contribute to inheritance of traits between parents and their children or grandparents and their grandchildren.
Another difference to ordinary mutations is that epimutations do not occur randomly, but often at the same places in the genome. While genes were disproportionately often affected, methylation of mobile DNA elements, transposons, was very stable. This appears to make sense, since it was already known that artificially induced loss of methylation has a much greater effect on the activity of transposons than of regular genes.

