Major advance in super-resolution fluorescence microscopy
Pushing the MINFLUX technique to higher spatial and temporal precision allows protein dynamics to be observed under physiological conditions
Scientists led by Nobel Laureate Stefan Hell at the Max Planck Institute for Medical Research in Heidelberg have developed a super-resolution microscope with a spatio-temporal precision of one nanometer per millisecond. An improved version of their recently introduced MINFLUX super-resolution microscopy allowed tiny movements of single proteins to be observed at an unprecedented level of detail: the stepping motion of the motor protein kinesin-1 as it walks along microtubules while consuming ATP. The work highlights the power of MINFLUX as a revolutionary new tool for observing nanometer-sized conformational changes in proteins.

Unraveling the inner workings of a cell requires knowledge of the biochemistry of individual proteins. Measuring tiny changes in their position and shape is the central challenge here. Fluorescence microscopy, in particular super-resolution microscopy (i.e. nanoscopy) has become indispensable in this emerging field. MINFLUX, the recently introduced fluorescence nanoscopy system, has already attained a spatial resolution of one to a few nanometers: the size of small organic molecules. But taking our understanding of molecular cell physiology to the next level requires observations at even higher spatio-temporal resolution.
When Stefan Hell’s group first presented MINFLUX in 2016, it had been used to track fluorescently labeled proteins in cells. However, these movements were random, and the tracking had precisions of the order of tens of nanometers. Their study is the first to apply the resolving power of MINFLUX to conformational changes of proteins, specifically the motor protein kinesin-1. To do this, the researchers at the Max Planck Institute for Medical Research developed a new MINFLUX version for tracking single fluorescent molecules.
All established methods for measuring protein dynamics have severe limitations, hampering their ability to address the critically important (sub)nanometer / (sub)millisecond range. Some provide a high spatial resolution, down to a few nanometers, but cannot track changes fast enough. Others have a high temporal resolution but require labeling with beads that are 2 to 3 orders of magnitude larger than the protein being studied. Since the functioning of the protein is likely to be compromised by a bead of this size, studies using beads leave open questions.
Fluorescence from a single molecule
MINFLUX, however, requires only a standard 1-nm sized fluorescence molecule as a label attached to the protein, and therefore can provide both the resolution and the minimal invasiveness that are needed in studying native protein dynamics. “One challenge lies in building a MINFLUX microscope that works close to the theoretical limit and is shielded against environmental noise”, says Otto Wolff, PhD student in the group. “Designing probes that do not affect the protein function, but still reveal the biological mechanism, is another”, adds his colleague Lukas Scheiderer.
The MINFLUX microscope which the researchers now introduce can record protein movements with a spatiotemporal precision of up to 1.7 nanometers per millisecond. It requires the detection of only about 20 photons emitted by the fluorescent molecule. “I think we are opening a new chapter in the study of the dynamics of individual proteins and how they change shape during their functioning”, says Stefan Hell. “The combination of high spatial and temporal resolution provided by MINFLUX will allow researchers to study biomolecules as never before.”
Resolving the stepping motion of kinesin-1 with ATP under physiological conditions
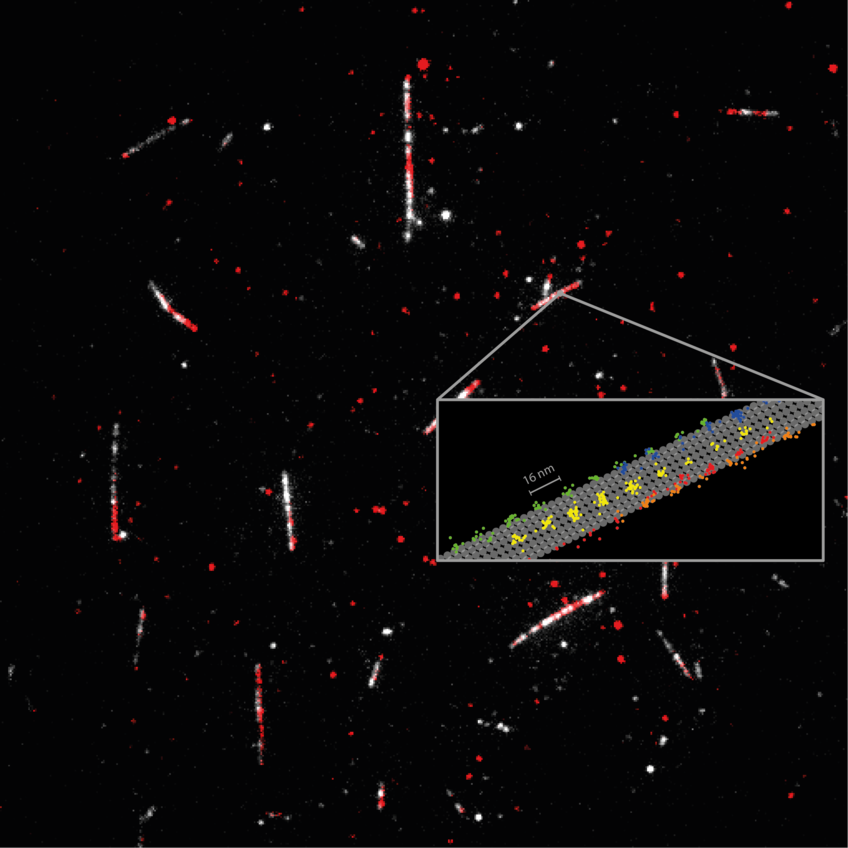
Kinesin-1 is a key player in transporting cargo throughout our cells, and mutations of the protein are at the heart of several diseases. Kinesin-1 actually ‘walks’ along filaments (the microtubules) that span our cells like a network of streets. One can imagine the motion as literally ’stepping‘, since the protein has two ‘heads’ that alternately change their location on the microtubule. This movement occurs usually along one of the 13 protofilaments forming the microtubule, and is fueled by splitting of the cell’s principal energy supplier ATP (adenosine triphosphate).
Using only a single fluorophore for labeling the kinesin-1, the scientists recorded the regular 16 nm. steps of individual heads as well as 8 nm substeps, with nanometer/millisecond spatiotemporal resolution. Their results proved that ATP is taken up while a single head is bound to the microtubule, but that ATP hydrolysis occurs when both heads are bound. It also revealed that the stepping involves a rotation of the protein ‘stalk’, the part of the kinesin molecule that holds the cargo. The spatiotemporal resolution of MINFLUX also revealed a rotation of the head in the initial phase of each step. Significantly, these findings were made using physiological concentrations of ATP, as was hitherto not possible with tiny fluorescence labels.
Future potential in exploring protein dynamics
“I’m excited so see where MINFLUX will take us. It adds another dimension to the study of how proteins work. This can help us to understand the mechanisms behind many diseases and ultimately contribute to the development of therapies”, adds Jessica Matthias, a postdoctoral scientist formerly in Hell’s group who is now exploring the applications of MINFLUX to a variety of biological questions.