A unique enzyme helps bacteria to breathe inside humans
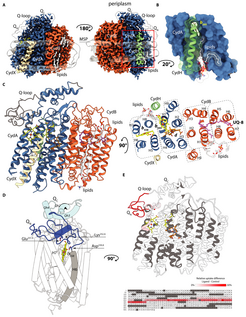
Structure of the cytochrome bd-I oxidase
Microorganisms have evolved a versatile repertoire of proteins and enzymes to survive under hostile environmental conditions. Cytochrome bd oxidases are membrane-integrated terminal oxygen reductases in branched respiratory chains of almost all bacteria. Their activity is essential for survival of commensal and pathogenic bacteria at low oxygen concentrations. An international team of scientists led by Hartmut Michel and Werner Kühlbrandt at the Max Planck Institute of Biophysics in Frankfurt now succeeded in determining the cryogenic electron microscopy structure of the prototype cytochrome bd oxidase from Escherichia coli.
In contrast to the mitochondrial type cytochrome c oxidases, the quinol oxidizing bd oxidases are insensitive to a broad range of toxic ligand compounds such as nitrogen monoxide (NO) released by macrophages or H2S enriched in the human gut. Due to its central role for survival of microorganisms in humans, the cytochrome bd oxidase is a target site for the development of next generation anti-microbial drugs to treat infectious diseases such as tuberculosis.
The study reveals structural divergence between the previously reported homolog from Geobacillus thermodenitfiricans (G. th) and the E. coli bd-I oxidase. Rearrangement of the active site heme groups and thereof resulting conformational adaptations propose an alternative oxygen binding site between the low-turnover enzyme from G.th and the high-turnover bd-I oxidase from E. coli.
Further, the research group identified a structural quinone molecule in the catalytically inactive CydB subunit and a proteobacteria specific accessory single transmembrane helix subunit bound to the catalytic CydA subunit. These findings suggest that assembly of the functional bd oxidase requires a higher-degree of regulation than previously assumed and may differ between oxidases from different phylae.