Adaptation and speciation mechanisms in sticklebacks
Yearbook article 2015 from the Friedrich Miescher Laboratory of the Max Planck Society
Organisms across the world show unique adaptations that enable them to survive and flourish in distinct environments. Researchers at the Friedrich Miescher Laboratory are studying stickleback fish to unravel the genetic changes which allow organisms to adapt and speciate in new environments. Marine sticklebacks have undergone an adaptive radiation with freshwater forms evolving repeatedly and independently at many different places. Using these powerful replicates of the evolutionary process, research is identifying the common molecular changes underlying adaptation and speciation.
Speciation
Earth is home to a remarkable variety of organisms, which come in all shapes and sizes. Many species have developed special traits to ensure their survival and reproduction in a particular environment. The study of the genetic basis and molecular mechanisms of such environmental adaptation may highlight the factors that reduce or promote fitness and survival in particular environments and therefore trigger the formation of new species. The results improve our understanding of the origins and development of biodiversity. Moreover, they provide clues as to how species, particularly agriculturally vital crops and animals, could adapt to rapid environmental change in the future. Finally, the results shed light on evolutionary fitness in different habitats and on the molecular basis of diseases caused by environmental factors.

Fig 1: The threespine stickleback (Gasterosteus aculeatus)
Stickleback fish as a model organism for the study of adaptive evolution
The threespine stickleback (Gasterosteus aculeatus, fig. 1) is a small marine and freshwater fish measuring 4 to 6 cm, which can be found in different aquatic habitats in temperate climate zones of the Northern hemisphere. Following the retreat of the last Pleistocene ice sheet between 10,000 and 20,000 years ago, ancestral marine sticklebacks colonized many newly formed freshwater habitats.
In a timespan that - in evolutionary terms - amounts to a blink of an eye, this species has undergone adaptive radiation: individuals ventured into new environments and adapted to the conditions there. Today, sticklebacks exist in many different shapes and sizes (fig. 2, [1]). Phenotypic (outward) diversity, resulting from the genotype’s interaction with the environment, is so great that natural historians and taxonomists initially described these populations as separate species. Nowadays, due to their ability to interbreed and produce viable and healthy offspring, the fish are consistently designated as a single species by researchers.
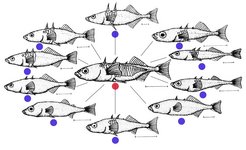
Fig. 2: In recent times, the threespine stickleback (Gasterosteus aculeatus) has undergone adaptive radiation, evolving from the typical marine ecotype (red) into freshwater ecotypes (blue) with a number of different shapes. Scale: 1 cm.
There are, however, remarkable exceptions with regard to the ability of the fish to interbreed and produce viable offspring – and that exactly what makes them so interesting to evolutionary biologists. One such exception – referred to as repeatedly evolved species pair – is distinct marine and freshwater stickleback forms, also called ecotypes. These morphologically, physiologically, behaviourally and genetically different ecotypes come into contact with each other during the breeding season, in the lower reaches of rivers and streams in the northern hemisphere [2]. Although hybridization and gene flow are possible and take place, the different ecotypes do not merge into a single homogenous population [3, 4]. On the contrary, the freshwater and marine fish retain their numerous traits in terms of shape, physiology and behaviour, and also display distinct gene pools. Studies confirm minor differences in mating preferences, and both hybrids and migrants can be observed in the contact zones [4, 5].
This contradicts the classical evolutionary theory, according to which the great gene flow would cause sticklebacks to form a new hybrid school. Researchers believe that hybrids and migrants are unable to survive or breed adequately if they possess the wrong genotype in the wrong habitat. Therefore heterogeneous marine and freshwater ecotypes are preserved. Since the phenomenon occurs repeatedly in different contact zones of marine and freshwater sticklebacks around the world, scientists now have a multitude of research sites at their disposal for more in-depth studies of the molecular basis of adaptation and speciation. Which factors and processes influence the diverse adaptation to different habitats consistently and generally? And how do they ultimately contribute to the formation of new species?
Sequencing of complete genomes reveals molecular basis of adaptation
Recently, a large-scale cooperation project employed the latest whole genome sequencing techniques to investigate the genomic basis for adaptive divergence between marine and freshwater sticklebacks [6]. An analysis of 21 marine and freshwater genomes, representing ten replicate pairs of marine and freshwater sticklebacks, offered the scientists in Tübingen excellent possibilities for exploring where and how evolution has optimized the genome.
The results were very interesting: In large parts of the genome, the freshwater sticklebacks did not differ from their marine neighbours. Such a pattern, caused only by random drift, is consistent with expectations based on evolutionary theory. However, geographically proximate marine and freshwater ecotypes displayed significant genetic variation in 81 different genome loci – corresponding to natural selection, which favours certain adaptive mutations in specific environments. In other words: the stickleback must have certain genetic variants in more than 81 different genomic loci in order to adapt to freshwater environments.
Reuse of existing genetic variation is important for adaptation to the habitat
The studies produced several surprising findings. Firstly, the genomic differences between marine and freshwater ecotypes are the result of parallel evolution during the process of radiation. For example, marine and freshwater sticklebacks in Scotland diverge in the same way as marine and freshwater fish in California with regard to the same genetic mutations in the same genome loci. The scientists assume that over 35% of the adaptive loci in the genome are cases of parallel reuse of a standing genetic variation (Fig. 3; [6]). These “prefabricated” adaptive building blocks can be found in all freshwater populations on Earth. The findings suggest that the building blocks are transported from one freshwater population to another through hybridization with proximate marine fish and are also introduced in new freshwater habitats by marine fish, with lower but verifiable frequency.
Individual marine ecotypes displaying freshwater-adaptive building blocks were discovered at a frequency of less than 2% [7]. Instead of waiting for suitable new mutations to occur – a process which can take millions of years – a population may in this way adapt rapidly to a new environment, as long as it possesses or has access to a useful standing genetic variation. The researchers are now studying thousands of marine sticklebacks to determine the factors and processes influencing the availability and preservation of adaptive genetic variability.
Adaptation using mutations in non-coding regions of the genome
The second surprising finding was that most of the mutations used for environmental adaptation apply to the non-coding regions of the genome ([6]; fig. 4). In comparison to the protein-coding genes, the non-coding DNA makes up the largest part of the genome, almost 90%. These regions of the DNA used to be considered junk DNA. However, thanks to scientific findings in the last 20 years, we now know that non-coding DNA contains important genetic switches, which regulate where and when a gene should be activated or deactivated in an organism.
The findings from the study of sticklebacks thus indicate that environmental adaptation mainly occurs through the mutation of non-coding elements that control the time and extent of gene and protein expression, rather than through alteration of the form or function of proteins themselves.
This discovery could affect the way scientists study the genetic basis of environmental diseases in humans. In fact, studies of all exons of an organism and all genome sequences which may code proteins could be overlooking crucial non-coding mutations that cause environmental diseases. The Tübingen-based laboratory is currently using a wide range of molecular biological methods to analyse the mechanisms that non-coding DNA elements use to regulate adaptive gene expression, which ultimately determine the fitness and survival of an individual.
Genomic inversion and adaptive gene cassettes
A third surprising discovery from the comparison of stickleback genomes was the complete inversion of three large gene regions between marine and freshwater ecotypes. These inversions are specifically able to keep the two inverted chromosome copies separate from one another by suppressing the combination of parental alleles in the gene transfer to offspring or, in other words: by suppressing recombination. This gives rise to a cassette of genes which is transferred intact and unaltered from one generation to the next. As unusual as this process may seem, it has proven a general evolutionary strategy for adaptation: sticklebacks, monkey flowers [8], apple maggot flies [9] and Heliconius butterflies [10] all display inversions in an early stage of adaptive divergence, as they have the ability to store several adaptive mutations as adaptive super-gene cassettes.
In order to continue to explore the genetic basis of variability with suppression of recombination, the author has recently received a Consolidator Grant of 2 million euros from the European Research Council (ERC).
The stickleback emerges as a super model organism for evolution which will boost progress in this research area
The modest threespine stickleback has turned out to be a super model organism for evolution, offering new insight into the molecular basis of adaptation and speciation. The adaptive radiation of this small fish provides numerous biological replicates and essential insight into molecular changes that enable organisms to colonise new environments quickly and to rapidly adapt to and survive in their new habitat. The research team is enthusiastically working on adding to the ever-growing stickleback genetic toolkit, which they can use to continuously gain new knowledge. Scientists can now use genome-editing techniques to investigate how specific adaptive mutations influence the phenotype and fitness of an individual. Next-generation sequencing technologies are employed to identify the recombination of hot and cold spots in the whole genome. Naturally, hybridizing populations are also studied to gain a better understanding of the way specific mutations affect hybrid fitness and limit gene flow between emerging marine and freshwater species.