Protein folding – why speed matters
Yearbook article 2016, Max Planck Institute for Molecular Biomedicine Author: Sebastian A. M. Leidel
Proteins are the workhorses of our cells. To fulfil their function, they need to adopt the correct shape. Scientists have now experimentally determined how fast proteins are made and have shown that the right speed is critical for functional folding. Abnormal translation rates lead to protein aggregates, which can cause severe developmental defects in mice, because cells in the brain receive the wrong differentiation signal due to protein stress. These results answer a fundamental question of molecular biology and have far-reaching consequences for neurodegenerative diseases and biotechnology.
How are the proteins in a cell produced? To answer this question, imagine the production line of a modern car factory. Various machines carry out a large number of steps in a defined order and in a precise rhythm. Only by means of this perfect interaction can a faultless car emerge at the end that will continue running for many years. Proteins, the workhorses of all cells, are produced in the human body in a similar manner. Ribosomes, large complexes of proteins and ribonucleic acid (RNA), link individual amino acids to form peptide chains (Fig. 1). These are folded into the right shape by special helpers called chaperones. But what happens when these processes go out of sync in cells? In a car factory, the answer is simple: if the production line were to slow down while the machines continued to operate at the same pace, welds would be made in the wrong locations, and the car would fail the final quality check.
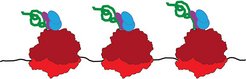
Fig. 1: Diagram of protein synthesis
Ribosomes (red) read messenger ribonucleic acid (black). The resulting amino acid chain (green) is folded into the correct conformation by chaperones (blue and purple).
In cells, the question is harder to address and has long puzzled scientists in the field of protein quality control. It was clear that chaperones are instrumental in ensuring that proteins assume their correct final shape. But what happens when the processing speed of ribosomes is disrupted? Importantly, this is not a purely academic question. When proteins develop quality issues, they clump together to form aggregates. Proteins in such aggregates can no longer function properly; indeed, they may even be toxic. This is very harmful for our cells, as can be seen by the fact that protein aggregates are implicated as the cause or symptom in a number of degenerative diseases. But does the function of proteins really depend on how fast they’re produced? As so often occurs in research, the answer to this question came from a completely different research area: the quality control of ribonucleic acids.
The curlicues of the genetic code
One often reads in textbooks that RNA molecules encode genetic information in four different building blocks, known as nucleotides. However, that does not do justice to the complexity of many naturally occurring RNA molecules. In fact, individual nucleotides of every RNA molecule in the cell are chemically modified (Fig. 2A) [1]. The modifications can be simple methyl groups, for example, but may also be composed of complex chemical groups (Fig. 2B).
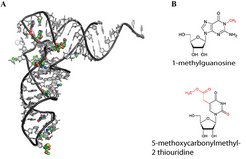
Fig. 2: Modified ribonucleic acid (RNA)
(A) Three-dimensional drawing of the structure of transfer RNA. Canonical nucleotides are shown in gray, chemical modifications as coloured spheres. (B) Two examples of RNA modifications: 1-methylguanosine has a simple methyl group. 5-methoxycarbonylmethyl-2 thiouridine is a complex modification and carries two different modifications. The modifications are shown in red.
It could be said that the genetic code has curlicues. And just as we ignore flourishes when reading text, many scientists ignored these RNA curlicues because their function was unclear. Yet there was evidence that the modifications are significant. Firstly, they occur in all organisms. Secondly, they often occur in functionally important sections of RNA molecules. Finally, it has been discovered in recent years that modification enzymes are mutated in a number of disorders, including neurodegenerative diseases, cancer and diabetes, to name only the most important conditions [2] − a good reason to take a closer look. And that’s exactly what the Max Planck Research Group for RNA Biology has done.
Rate control for protein synthesis
The working hypothesis was that the absence of modifications affects the rate of protein production. But how can this premise be tested? In the case of a car, you could stand at the side of the road and measure the car’s speed. But that’s not so easy with ribosomes. Technical progress, however, has come to the researchers’ aid in the form of a technique known as ribosome profiling (Fig. 3) [3].
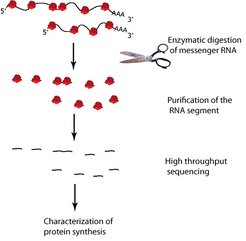
Fig. 3: Diagram of ribosome profiling
Ribosomes (red) read messenger RNA (black). Enzymes digest any messenger RNA outside the ribosomes. The RNA segments are then separated from the ribosomes, sequenced and characterized.
This new method makes it possible to determine which proteins are being manufactured by the ribosomes of a cell. The really fascinating thing about the method is that it allows to analyse many details of the reading process, including its rate. To this end, during ribosome profiling by high-throughput sequencing, all RNA segments that are currently being translated into a protein by ribosomes are read. If enough segments are observed, a conclusion can be drawn regarding all the ribosomes of a cell. However, it takes about three weeks for the scientists to save the many millions of sequence segments of an experiment in their computer. And then the sequences have to be read and interpreted. Ultimately, though, the ribosomes reveal their processing speed, as if the experimenters had placed speed cameras in the cells. Yeast cells have proved suitable for such experiments, as they are simply constructed, and comprise the same components as a human cell: the cell nucleus stores genetic information, mitochondria generate energy and ribosomes act as protein production lines. Moreover, yeast mutants in which modification enzymes are defective, are easily obtained.
Who brakes, loses!
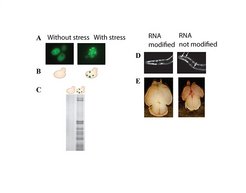
Fig. 4: Stress due to protein aggregates in cells with slowed protein synthesis (modification mutants).
(A) fluorescence micrograph of a yeast modification mutant. Protein aggregates are visualized by means of a reporter protein. (B) Drawing of (A). Protein aggregates are shown as red dots, reporter protein as green dots. (C) Biochemical isolation of protein aggregates. Protein aggregates from yeast cells appear as brown bands in a protein gel. Only the modification mutant (right) contains aggregates, while normal yeast (left) contains none. Right: morphological phenotypes in mutants for RNA modification enzymes in (D) nematodes and (E) mice. (D) Fluorescence micrographs of nematodes show a diffuse protein signal (white arrow). (E) Mouse brains shortly after birth. The brain of a mutant is significantly smaller than normal (bottom).
The data immediately showed that ribosomes are still able to read the genetic information even if the RNA is unmodified. Only those sections, or “words”, that require RNA modifications are read much more slowly in modification mutants than in normal yeast cells [4]. By contrast, the accuracy of reading appeared to be unaffected. This was the first direct indication that ribosomes actually read more slowly when RNA modifications are absent. The opportunity presented by these mutants was immediately clear: previously, scientists were only able to alter the overall rate of protein production with the help of inhibitors. With modification mutants, it is now possible to study what happens when only individual parts of the production process are disrupted while the other processes apparently continue to operate normally.
What happens when ribosome coordination is lost? The cells fill up with protein aggregates. That was unexpected, and it was therefore important to look at the mechanisms by which cells protects themselves from the harmful effects of aggregates. Cells try either to repair aggregated proteins with the help of chaperones or to degrade them with the help of their disposal system. The experiments showed that both systems were completely overwhelmed (Fig. 4A). In this way, it was possible to isolate the protein aggregates biochemically from the cells. Protein aggregates do not occur when the normal speed is restored in the slow sections of RNA. Speed must therefore be crucial. Damage results if an imbalance occurs in the interplay between the production rate of proteins and folding by chaperones. The ”wisdom” of bumper stickers applies here: Who brakes, loses!
Whether yeast, worms or mice - the problem remains the same
The next surprise came when the researchers looked at the proteins contained in the aggregates. They found that the aggregated proteins are not those that are read slowly, but those that are unstable within the cell. Even in normal situations, cells need key enzymes that are hardly soluble and therefore tend to aggregate. The fact that the reading speed is disturbed in mutants means that the cell’s quality control system is no longer able to repair these difficult proteins, so that they clump together. Consequently, the cell fails as a system. This was an entirely new approach to explaining the defects of modification mutants.
Can these findings in yeast be transferred to other organisms? This is an important question if we want to shed light on diseases. An answer was obtained in two steps: Nematodes, which possess a nervous system, are easy to study. Under the microscope it can readily be seen that modification mutants contain protein aggregates and do not live as long as normal nematodes (Fig. 4B) [4].
But are similar mechanisms at work in vertebrates? The scientists sought to answer this question using the example of a rare genetic disease. In familial dysautonomia, a modification enzyme is defective, which means that part of the nervous system is not created. As a result, affected individuals are barely able to perceive sensory stimuli and die at a relatively young age. To investigate whether this is related to the findings in yeast and worms, the Research Group worked in cooperation with the laboratory of Laurent Nguyen [5]. The Belgian researchers had mice in whose brain the enzyme is dysfunctional. Again, it turned out that protein production is slowed, resulting in protein stress. The upshot is that the mice are born with a smaller brain than their healthy siblings (Fig. 4D). Thus, an apparently small effect has enormous consequences.
It has been established that protein production is a complex process in which many factors play a role. The Max Planck Research Group for RNA Biology has now demonstrated just how critical the production rate is, and this aspect has to be borne in mind in the future. In the past, researchers generally ignored the production speed of proteins and were only interested in their folding and degradation. This must now change. The topic also impacts biotechnology. The methods used for synthesizing proteins in the chemical and pharmaceutical industries are often very difficult and inefficient. A close look at the production speed may also work wonders here.



