Gene knockout: loss of a gene can be compensated by another gene
Effects of genome interventions depend on the methods used
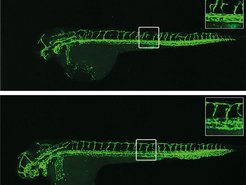
Blood vessels of a zebrafish larva: if the gene EGFL7 is lacking, the blood vessels (stained green) are not formed correctly. In genetically unchanged larvae which possess the EGFL7 gene, a substance that blocks EGFL7 leads to disturbances in vessel growth (above). In fish larvae without the EGFL7 gene, however, other genes compensate for the loss, so that the inhibitor hardly affects the growth of blood vessels (below).
To analyse function of an unkown gene, scientists often extinguish the gene and investigate the consequences of this treatment for the organism. To do so, they cut DNA-fragments from the gene using enzymes deleting the genetic information for a functioning protein. Such method is called “Gene knockout”. In contrast, in a “gene knockdown” scientists block protein production using particular substances, e.g. microRNAs.
Recent studies, however, have shown that results may vary between knockout- and knockdown animals. Scientists from Didier Stainier's group at the Max Planck Institute for Heart and Lung Research have now identified the reason for this. The Bad Nauheim based researchers have investigated a gene called egfl7 in zebrafish. The gene is involved in the production of connective tissue in blood vessel walls, thereby stabilizing them. Doing so, egfl7 regulates blood vessel growth.
Developmental biologists, however, are not sure, what happens in a fish organism, after the egfl7 gene has been deleted. “If the gene has been blocked in a knockdown, blood vessels do not develop normally”, explains Andrea Rossi, together with Zacharias Kontarakis first author of the study. In contrast, if the gene itself is deleted by a genetic manipulation, blood vessel growth is not affected.
In the beginning, Max Planck researchers excluded potential side effects of the knockdown substance being responsible for interference in vascular development. To this end, they injected the substance into fish larvae in which the egfl7 gene had already been deleted. However, the larvae almost developed normally.
“Since the substance did not cause disturbances in blood vessel growth, we thought of a different mechanism: The gene loss could be compensated by another gene taking over the function”, Kontarakis says. “Therefore, we were looking for rescue genes, which might have been produced in animals without a functional egfl7 gene.”
The researchers compared the mRNA molecules and proteins in fish with or without a functional egfl7 gene and detected several mRNAs and proteins being present in higher amounts in fish without egfl7. An example is emilin 3B. When “knockdown” animals are treated with emilin 3B after egfl7 has been blocked, blood vessels develop almost normally. “This tells us that emilin 3B can compensate for the loss of egfl7. In egfl7 knockout fish, emilin production is getting upregulated. This is not the case in knockdown fish”, Stainier explains.
As the next step, the group plans to analyse how genes “know” that another gene has been deleted and then compensate for the loss. Several researchers worldwide are trying to delete disease genes for therapeutic reasons. Before we establish such therapies, we have to fully understand the consequences the loss or blockade of a gene might have. “In addition, our study illustrates the power of comparing knockouts and knockdowns to identify modifier genes, a goal that remains a major challenge in the field of human genetics” says Stainier.
MH/HR
